TheiaCoV Workflow Series¶
The TheiaCoV Workflow Series is a collection of WDL workflows developed for performing genomic characterization and genomic epidemiology of SARS-CoV-2 samples to support public health decision-making.
TheiaCoV Workflows for Genomic Characterization¶
Genomic characterization, i.e. generating consensus assemblies (FASTA format) from next-generation sequencing (NGS) read data (FASTQ format) to assign samples with relevant nomenclature designation (e.g. PANGO lineage and NextClade clades) is an increasingly critical function to public health laboratories around the world.
The TheiaCoV Genomic Characterization Series includes four separate WDL workflows (TheiaCoV_Illumina_PE, TheiaCoV_Illumina_SE, TheiaCoV_ClearLabs, and TheiaCoV_ONT) that process NGS read data from four different sequencing approaches: Illumina paired-end, Illumina single-end, Clear Labs, and Oxford Nanopore Technology (ONT)) to generate consensus assemblies, produce relevant quality-control metrics for both the input read data and the generated assembly, and assign samples with a lineage and clade designation using Pangolin and NextClade, respectively.
All four TheiaCoV workflows for genomic characterization will generate a viral assembly by mapping input read data to a reference genome, removing primer reads from that alignment, and then calling the consensus assembly based on the primer-trimmed alignment. These consensus assemblies are then fed into the Pangolin and NextClade CLI tools for lineage and clade assignments.
The major difference between each of these TheiaCoV Genomic Characterization workflows is in how the read mapping, primer trimming, and consensus genome calling is performed. More information on the technical details of these processes and information on how to utilize and apply these workflows for public health investigations is available below.
A fifth WDL workflow, TheiaCoV_FASTA, was added to take in assembled SC2 genomes, perform basic QC (e.g. number of Ns), and assign samples with a lineage and clade designation using Pangolin and NextClade, respectively.
A series of introductory training videos that provide conceptual overviews of methodologies and walkthrough tutorials on how to utilize these TheiaCoV workflows through Terra are available on the Theiagen Genomics YouTube page:
note Titan workflows in the video have since been renamed to TheiaCoV.
TheiaCoV_Illumina_PE¶
The TheiaCoV_Illumina_PE workflow was written to process Illumina paired-end (PE) read data. Input reads are assumed to be the product of sequencing tiled PCR-amplicons designed for the SARS-CoV-2 genome. The most common read data analyzed by the TheiaCoV_Illumina_PE workflow are generated with the Artic V3 protocol. Alternative primer schemes such as the Qiaseq Primer Panel, the Swift Amplicon SARS-CoV-2 Panel and the Artic V4 Amplicon Sequencing Panel however, can also be analysed with this workflow since the primer sequence coordinates of the PCR scheme utilized must be provided along with the raw paired-end Illumina read data in BED and FASTQ file formats, respectively.
Note
By default, this workflow will assume that input reads were generated using a 300-cycle kit (i.e. 2 x 150 bp reads). Modifications to the optional parameter for trimmomatic_minlen may be required to accommodate for shorter read data, such as 2 x 75bp reads generated using a 150-cycle kit.
Upon initiating a TheiaCoV_Illumina_PE job, the input primer scheme coordinates and raw paired-end Illumina read data provided for each sample will be processed to perform consensus genome assembly, infer the quality of both raw read data and the generated consensus genome, and assign SARS-CoV-2 lineage and clade types as outlined in the TheiaCoV_Illumina_PE data workflow below.
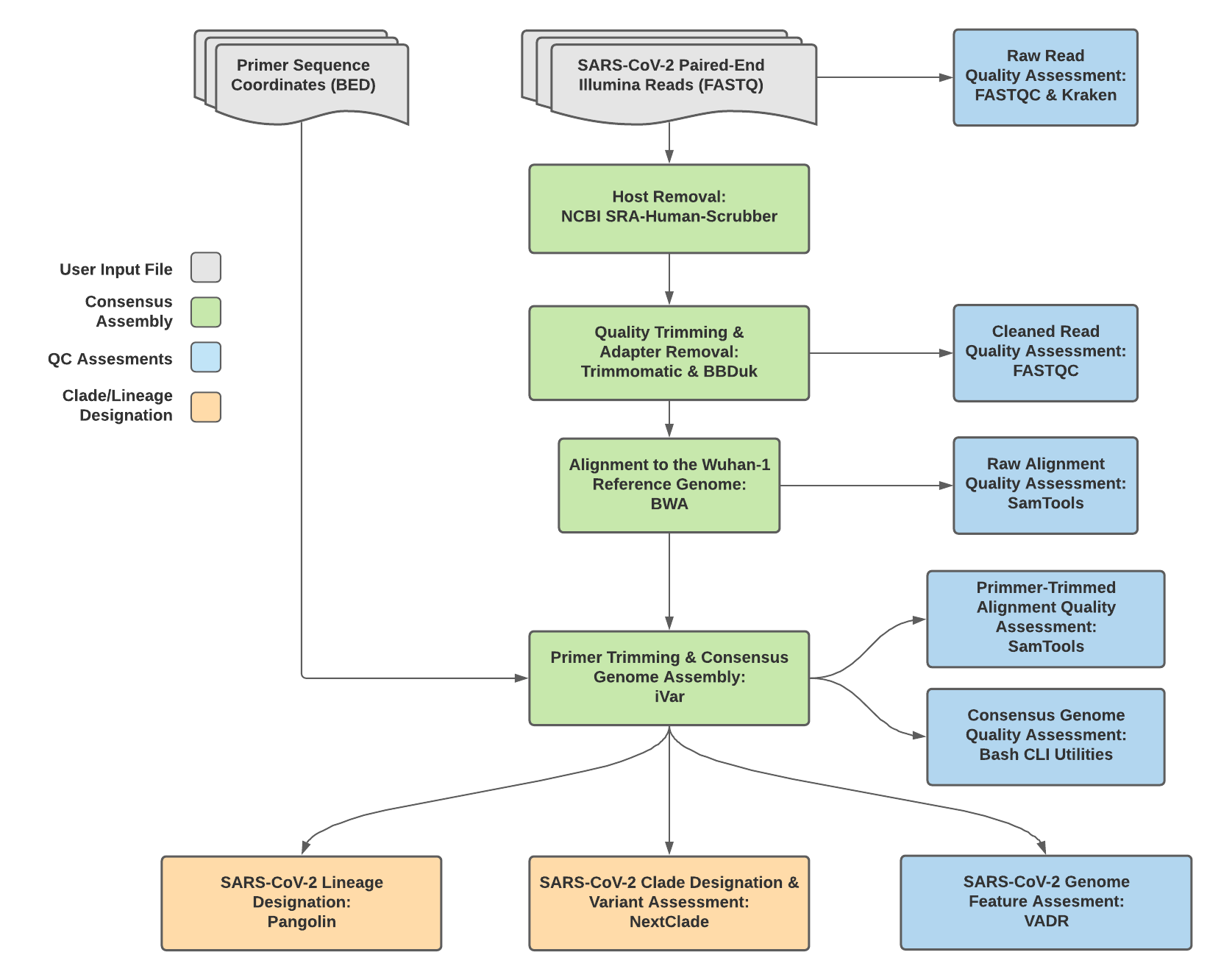
TheiaCoV_Illumina_PE Data Workflow¶
Consensus genome assembly with the TheiaCoV_Illumina_PE workflow is performed by first de-hosting read data with the NCBI SRA-Human-Scrubber tool then trimming low-quality reads with Trimmomatic and removing adapter sequences with BBDuk. These cleaned read data are then aligned to the Wuhan-1 reference genome with BWA to generate a Binary Alignment Mapping (BAM) file. Primer sequences are then removed from the BAM file using the iVar Trim sub-command. The iVar consensus sub-command is then utilized to generate a consensus assembly in FASTA format. This assembly is then used to assign lineage and clade designations with Pangolin and NextClade. NCBI’S VADR tool is also employed to screen for potentially errant features (e.g. erroneous frame-shift mutations) in the consensus assembly.
More information on required user inputs, optional user inputs, default tool parameters and the outputs generated by TheiaCoV_Illumina_PE are outlined below.
Required User Inputs¶
Download CSV: TheiaCoV_Illumina_PE_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
theiacov_illumina_pe |
primer_bed |
File |
Primer sequence coordinates of the PCR scheme utilized in BED file format |
theiacov_illumina_pe |
read1_raw |
File |
Forward Illumina read in FASTQ file format |
theiacov_illumina_pe |
read2_raw |
File |
Reverse Illumina read in FASTQ file format |
theiacov_illumina_pe |
samplename |
String |
Name of the sample being analyzed |
Optional User Inputs¶
Download CSV: TheiaCoV_Illumina_PE_optional_inputs.csv
Task |
Variable Name |
Data Type |
Description |
Default |
|---|---|---|---|---|
bwa |
reference_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
bwa |
cpus |
Int |
CPU resources allocated to the BWA task runtime environment |
6 |
consensus |
char_unknown |
String |
Character to print in regions with less than minimum coverage for iVar consensus |
N |
consensus |
count_orphans |
Boolean |
Do not skip anomalous read pairs in variant calling for SAMtools mpileup before running iVar consensus |
TRUE |
consensus |
disable_baq |
Boolean |
Disable read-pair overlap detection for SAMtools mpileup before running iVar consensus |
TRUE |
consensus |
max_depth |
Int |
Maximum reads read at a position per input file for SAMtools mpileup before running iVar consensus |
600000 |
consensus |
min_bq |
Int |
Minimum mapping quality for an alignment to be used for SAMtools mpileup before running iVar consensus |
0 |
consensus |
min_depth |
Int |
Minimum read depth to call variants for iVar consensus |
10 |
consensus |
min_freq |
Float |
Minimum frequency threshold(0 - 1) to call variants for iVar consensus |
0.6 |
consensus |
min_qual |
Int |
Minimum quality threshold for sliding window to pass for iVar consensus |
20 |
consensus |
ref_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
consensus |
ref_gff |
String |
Path to the general feature format of the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/reference/GCF_009858895.2_ASM985889v3_genomic.gff |
nextclade_one_sample |
docker |
String |
Docker tag used for running NextClade |
neherlab/nextclade:0.14.2 |
nextclade_output_parser_one_sample |
docker |
String |
Docker tag used for parsing NextClade output |
python:slim |
pangolin3 |
docker |
String |
Docker tag used for running Pangolin |
staphb/pangolin:3.1.11-pangolearn-2021-08-24 |
pangolin3 |
inference_engine |
String |
pangolin inference engine for lineage designations (usher or pangolarn) |
usher |
pangolin3 |
min_length |
Int |
Minimum query length allowed for pangolin to attempt assignment |
10000 |
pangolin3 |
max_ambig |
Float |
Maximum proportion of Ns allowed for pangolin to attempt assignment |
0.5 |
primer_trim |
keep_noprimer_reads |
Boolean |
Include reads with no primers for iVar trim |
True |
read_QC_trim |
bbduk_mem |
Int |
Memory allocated to the BBDuk VM |
8 |
read_QC_trim |
trimmomatic_minlen |
Int |
Specifies the minimum length of reads to be kept for Trimmomatic |
25 |
read_QC_trim |
trimmomatic_quality_trim_score |
Int |
Specifies the average quality required for Trimmomatic |
30 |
read_QC_trim |
trimmomatic_window_size |
Int |
Specifies the number of bases to average across for Trimmomatic |
4 |
theiacov_illumina_pe |
nextclade_dataset_name |
String |
Nextclade organism dataset |
sars-cov-2 |
theiacov_illumina_pe |
nextclade_dataset_reference |
String |
Nextclade reference genome |
MN908947 |
theiacov_illumina_pe |
nextclade_dataset_tag |
Nextclade dataset tag |
2021-06-25T00:00:00Z |
|
theiacov_illumina_pe |
seq_method |
String |
Description of the sequencing methodology used to generate the input read data |
Illumina paired-end |
vadr |
docker |
String |
Docker tag used for running VADR |
staphb/vadr:1.2.1 |
vadr |
maxlen |
Int |
Maximum length for the fasta-trim-terminal-ambigs.pl VADR script |
30000 |
vadr |
minlen |
Int |
Minimum length subsequence to possibly replace Ns for the fasta-trim-terminal-ambigs.pl VADR script |
50 |
vadr |
skip_length |
Int |
Minimum assembly length (unambiguous) to run vadr |
10000 |
vadr |
vadr_opts |
String |
Options for the v-annotate.pl VADR script |
–glsearch -s -r –nomisc –mkey sarscov2 –alt_fail lowscore,fstukcnf,insertnn,deletinn –mdir /opt/vadr/vadr-models/ |
variant_call |
count_orphans |
Boolean |
Do not skip anomalous read pairs in variant calling for SAMtools mpileup before running iVar variants |
TRUE |
variant_call |
disable_baq |
Boolean |
Disable read-pair overlap detection for SAMtools mpileup before running iVar variants |
TRUE |
variant_call |
max_depth |
Int |
Maximum reads read at a position per input file for SAMtools mpileup before running iVar variants |
600000 |
variant_call |
min_bq |
Int |
Minimum mapping quality for an alignment to be used for SAMtools mpileup before running iVar variants |
0 |
variant_call |
min_depth |
Int |
Minimum read depth to call variants for iVar variants |
10 |
variant_call |
min_freq |
Float |
Minimum frequency threshold(0 - 1) to call variants for iVar variants |
0.6 |
variant_call |
min_qual |
Int |
Minimum quality threshold for sliding window to pass for iVar variants |
20 |
variant_call |
ref_gff |
String |
Path to the general feature format of the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/reference/GCF_009858895.2_ASM985889v3_genomic.gff |
variant_call |
ref_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
version_capture |
timezone |
String |
User time zone in valid Unix TZ string (e.g. America/New_York) |
None |
Outputs¶
Download CSV: TheiaCoV_Illumina_PE_default_outputs.csv
Output Name |
Data Type |
Description |
|---|---|---|
aligned_bai |
File |
Index companion file to the bam file generated during the consensus assembly process |
aligned_bam |
File |
Primer-trimmed BAM file; generated during conensus assembly process |
assembly_fasta |
File |
Consensus genome assembly |
assembly_length_unambiguous |
Int |
Number of unambiguous basecalls within the SC2 consensus assembly |
assembly_mean_coverage |
Float |
Mean sequencing depth throughout the conesnsus assembly generated after performing primer trimming–calculated using the SAMtools coverage command |
assembly_method |
String |
Method employed to generate consensus assembly |
auspice_json |
File |
Auspice-compatable JSON output generated from NextClade analysis that includes the NextClade default samples for clade-typing and the single sample placed on this tree |
bbduk_docker |
String |
Docker image used to run BBDuk |
bwa_version |
String |
Version of BWA used to map read data to the reference genome |
consensus_flagstat |
File |
Output from the SAMtools flagstat command to assess quality of the alignment file (BAM) |
consensus_stats |
File |
Output from the SAMtools stats command to assess quality of the alignment file (BAM) |
fastqc_clean1 |
Int |
Number of forward reads after seqyclean filtering as determined by FastQC |
fastqc_clean2 |
Int |
Number of reverse reads after seqyclean filtering as determined by FastQC |
fastqc_clean_pairs |
String |
Number of paired reads after SeqyClean filtering as determined by FastQC |
fastqc_raw1 |
Int |
Number of forward reads identified in the input fastq files as determined by FastQC |
fastqc_raw2 |
Int |
Number of reverse reads identified in the input fastq files as determined by FastQC |
fastqc_raw_pairs |
String |
Number of paired reads identified in the input fastq files as determined by FastQC |
fastqc_version |
String |
Version of the FastQC software used for read QC analysis |
ivar_tsv |
File |
Variant descriptor file generated by iVar variants |
ivar_variant_version |
String |
Version of iVar for running the iVar variants command |
ivar_vcf |
File |
iVar tsv output converted to VCF format |
ivar_version_consensus |
String |
Version of iVar for running the iVar consensus command |
ivar_version_primtrim |
String |
Version of iVar for running the iVar trim command |
kraken_human |
Float |
Percent of human read data detected using the Kraken2 software |
kraken_human_dehosted |
Float |
Percent of human read data detected using the Kraken2 software after host removal |
kraken_report |
File |
Full Kraken report |
kraken_report_dehosted |
File |
Full Kraken report after host removal |
kraken_sc2 |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software |
kraken_sc2_dehosted |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software after host removal |
kraken_version |
String |
Version of Kraken software used |
meanbaseq_trim |
Float |
Mean quality of the nucleotide basecalls aligned to the reference genome after primer trimming |
meanmapq_trim |
Float |
Mean quality of the mapped reads to the reference genome after primer trimming |
nextclade_aa_dels |
String |
Amino-acid deletions as detected by NextClade |
nextclade_aa_subs |
String |
Amino-acid substitutions as detected by NextClade |
nextclade_clade |
String |
NextClade clade designation |
nextclade_json |
File |
NexClade output in JSON file format |
nextclade_tsv |
File |
NextClade output in TSV file format |
nextclade_version |
String |
Version of NextClade software used |
number_Degenerate |
Int |
Number of degenerate basecalls within the consensus assembly |
number_N |
Int |
Number of fully ambiguous basecalls within the consensus assembly |
number_Total |
Int |
Total number of nucleotides within the consensus assembly |
pango_lineage |
String |
Pango lineage as detremined by Pangolin |
pango_lineage_report |
File |
Full Pango lineage report generated by Pangolin |
pangolin_assignment_version |
String |
Version of the pangolin software (e.g. PANGO or PUSHER) used for lineage asignment |
pangolin_conflicts |
String |
Number of lineage conflicts as deteremed by Pangolin |
pangolin_docker |
String |
Docker image used to run Pangolin |
pangolin_notes |
String |
Lineage notes as deteremined by Pangolin |
pangolin_versions |
String |
All Pangolin software and database version |
percent_reference_coverage |
Float |
Percent coverage of the reference genome after performing primer trimming; calculated as assembly_length_unambiguous / length of reference genome (SC2: 29,903) x 100 |
primer_bed_name |
String |
Name of the primer bed files used for primer trimming |
primer_trimmed_read_percent |
Float |
Percent of read data with primers trimmed as deteremined by iVar trim |
read1_clean |
File |
Forward read file after quality trimming and adapter removal |
read1_dehosted |
File |
Dehosted forward reads; suggested read file for SRA submission |
read2_clean |
File |
Reverse read file after quality trimming and adapter removal |
read2_dehosted |
File |
Dehosted reverse reads; suggested read file for SRA submissionsamtools_version |
samtools_version |
String |
Version of SAMtools used to sort and index the alignment file |
samtools_version_consensus |
String |
Version of SAMtools used to create the pileup before running iVar consensus |
samtools_version_primtrim |
String |
Version of SAMtools used to create the pileup before running iVar trim |
samtools_version_stats |
String |
Version of SAMtools used to assess quality of read mapping |
seq_platform |
String |
Description of the sequencing methodology used to generate the input read data |
theiacov_illumina_pe_analysis_date |
String |
Date of analysis |
theiacov_illumina_pe_version |
String |
Version of the Public Health Viral Genomics (PHVG) repository used |
trimmomatic_version |
String |
Version of Trimmomatic used |
vadr_alerts_list |
File |
File containing all of the fatal alerts as determined by VADR |
vadr_docker |
String |
Docker image used to run VADR |
vadr_num_alerts |
String |
Number of fatal alerts as determined by VADR |
TheiaCoV_Illumina_SE¶
The TheiaCoV_Illumina_SE workflow was written to process Illumina single-end (SE) read data. Input reads are assumed to be the product of sequencing tiled PCR-amplicons designed for the SARS-CoV-2 genome. The most common read data analyzed by the TheiaCoV_Illumina_SE workflow are generated with the Artic V3 protocol. Alternative primer schemes such as the Qiaseq Primer Panel, however, can also be analysed with this workflow since the primer sequence coordinates of the PCR scheme utilized must be provided along with the raw paired-end Illumina read data in BED and FASTQ file formats, respectively.
Note
By default, this workflow will assume that input reads were generated using a 35-cycle kit (i.e. 1 x 35 bp reads). Modifications to the optional parameter for trimmomatic_minlen may be required to accommodate for longer read data.
Upon initiating a TheiaCoV_Illumina_SE job, the input primer scheme coordinates and raw paired-end Illumina read data provided for each sample will be processed to perform consensus genome assembly, infer the quality of both raw read data and the generated consensus genome, and assign SARS-CoV-2 lineage and clade types as outlined in the TheiaCoV_Illumina_PE data workflow below.
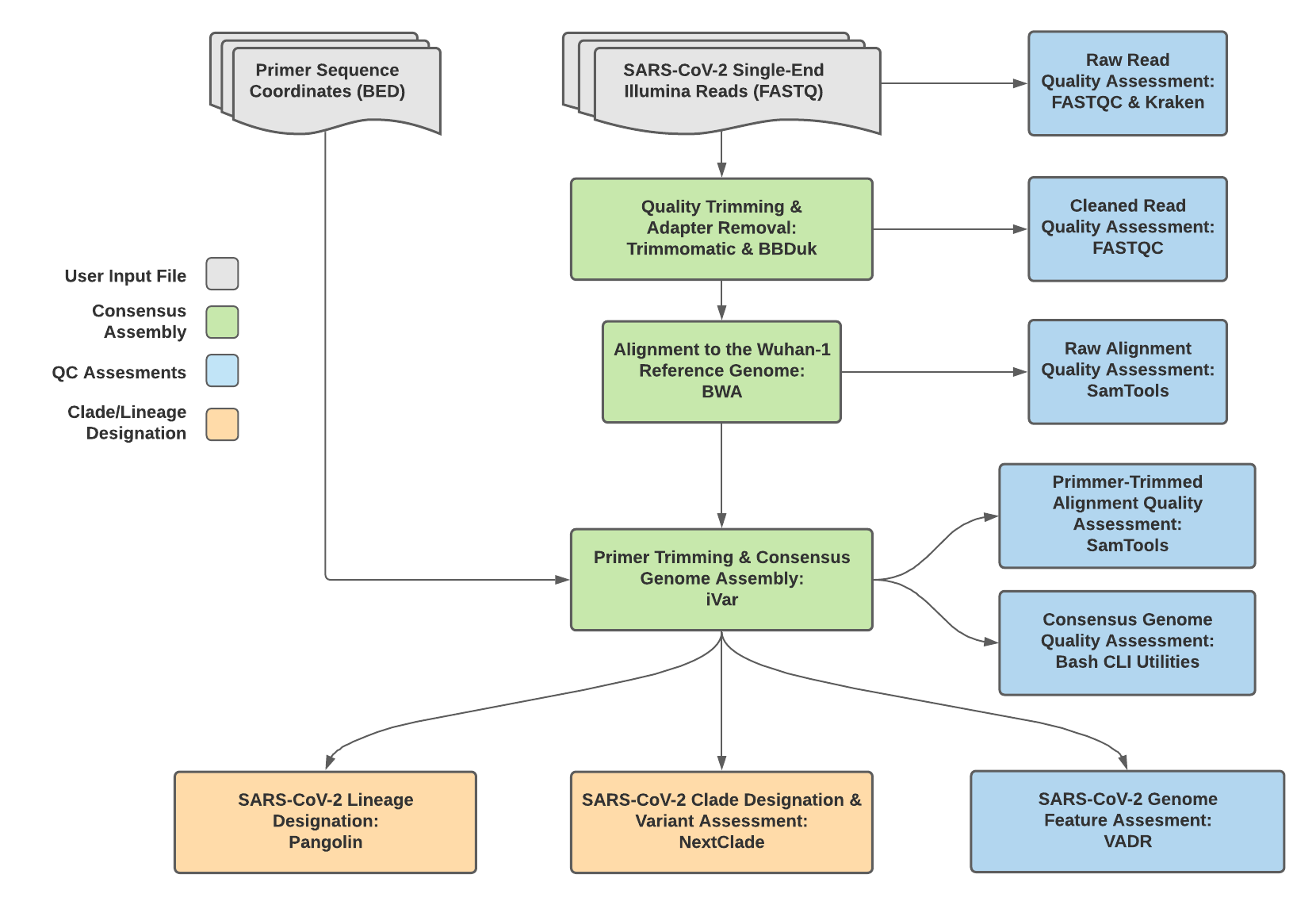
TheiaCoV_Illumina_SE Data Workflow¶
Consensus genome assembly with the TheiaCoV_Illumina_SE workflow is performed by first trimming low-quality reads with Trimmomatic and removing adapter sequences with BBDuk. These cleaned read data are then aligned to the Wuhan-1 reference genome with BWA to generate a Binary Alignment Mapping (BAM) file. Primer sequences are then removed from the BAM file using the iVar Trim sub-command. The iVar consensus sub-command is then utilized to generate a consensus assembly in FASTA format. This assembly is then used to assign lineage and clade designations with Pangolin and NextClade. NCBI’S VADR tool is also employed to screen for potentially errant features (e.g. erroneous frame-shift mutations) in the consensus assembly.
More information on required user inputs, optional user inputs, default tool parameters and the outputs generated by TheiaCoV_Illumina_SE are outlined below.
Required User Inputs¶
Download CSV: TheiaCoV_Illumina_SE_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
theiacov_illumina_pe |
primer_bed |
File |
Primer sequence coordinates of the PCR scheme utilized in BED file format |
theiacov_illumina_pe |
read1_raw |
File |
Single-end Illumina read in FASTQ file format |
theiacov_illumina_pe |
samplename |
String |
Name of the sample being analyzed |
Optional User Inputs¶
Download CSV: TheiaCoV_Illumina_SE_optional_inputs.csv
Task |
Variable Name |
Data Type |
Description |
Default |
|---|---|---|---|---|
bwa |
reference_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
bwa |
cpus |
Int |
CPU resources allocated to the BWA task runtime environment |
6 |
bwa |
read2 |
File |
Optional input file for the Kraken task that is not applicable to this workflow |
None |
consensus |
char_unknown |
String |
Character to print in regions with less than minimum coverage for iVar consensus |
N |
consensus |
count_orphans |
Boolean |
Do not skip anomalous read pairs in variant calling for SAMtools mpileup before running iVar consensus |
TRUE |
consensus |
disable_baq |
Boolean |
Disable read-pair overlap detection for SAMtools mpileup before running iVar consensus |
TRUE |
consensus |
max_depth |
Int |
Maximum reads read at a position per input file for SAMtools mpileup before running iVar consensus |
600000 |
consensus |
min_bq |
Int |
Minimum mapping quality for an alignment to be used for SAMtools mpileup before running iVar consensus |
0 |
consensus |
min_depth |
Int |
Minimum read depth to call variants for iVar consensus |
10 |
consensus |
min_freq |
Float |
Minimum frequency threshold(0 - 1) to call variants for iVar consensus |
0.6 |
consensus |
min_qual |
Int |
Minimum quality threshold for sliding window to pass for iVar consensus |
20 |
consensus |
ref_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
consensus |
ref_gff |
String |
Path to the general feature format of the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/reference/GCF_009858895.2_ASM985889v3_genomic.gff |
nextclade_one_sample |
docker |
String |
Docker tag used for running NextClade |
neherlab/nextclade:0.14.2 |
nextclade_output_parser_one_sample |
docker |
String |
Docker tag used for parsing NextClade output |
python:slim |
pangolin3 |
docker |
String |
Docker tag used for running Pangolin |
staphb/pangolin:3.1.11-pangolearn-2021-08-24 |
pangolin3 |
inference_engine |
String |
pangolin inference engine for lineage designations (usher or pangolarn) |
usher |
pangolin3 |
min_length |
Int |
Minimum query length allowed for pangolin to attempt assignment |
10000 |
pangolin3 |
max_ambig |
Float |
Maximum proportion of Ns allowed for pangolin to attempt assignment |
0.5 |
primer_trim |
keep_noprimer_reads |
Boolean |
Include reads with no primers for iVar trim |
True |
read_QC_trim |
bbduk_mem |
Int |
Memory allocated to the BBDuk VM |
8 |
read_QC_trim |
trimmomatic_minlen |
Int |
Specifies the minimum length of reads to be kept for Trimmomatic |
25 |
read_QC_trim |
trimmomatic_quality_trim_score |
Int |
Specifies the average quality required for Trimmomatic |
30 |
read_QC_trim |
trimmomatic_window_size |
Int |
Specifies the number of bases to average across for Trimmomatic |
4 |
theiacov_illumina_se |
nextclade_dataset_name |
String |
Nextclade organism dataset |
sars-cov-2 |
theiacov_illumina_se |
nextclade_dataset_reference |
String |
Nextclade reference genome |
MN908947 |
theiacov_illumina_se |
nextclade_dataset_tag |
Nextclade dataset tag |
2021-06-25T00:00:00Z |
|
theiacov_illumina_se |
seq_method |
String |
Description of the sequencing methodology used to generate the input read data |
Illumina paired-end |
vadr |
docker |
String |
Docker tag used for running VADR |
staphb/vadr:1.2.1 |
vadr |
maxlen |
Int |
Maximum length for the fasta-trim-terminal-ambigs.pl VADR script |
30000 |
vadr |
minlen |
Int |
Minimum length subsequence to possibly replace Ns for the fasta-trim-terminal-ambigs.pl VADR script |
50 |
vadr |
skip_length |
Int |
Minimum assembly length (unambiguous) to run vadr |
10000 |
vadr |
vadr_opts |
String |
Options for the v-annotate.pl VADR script |
–glsearch -s -r –nomisc –mkey sarscov2 –alt_fail lowscore,fstukcnf,insertnn,deletinn –mdir /opt/vadr/vadr-models/ |
variant_call |
count_orphans |
Boolean |
Do not skip anomalous read pairs in variant calling for SAMtools mpileup before running iVar variants |
TRUE |
variant_call |
disable_baq |
Boolean |
Disable read-pair overlap detection for SAMtools mpileup before running iVar variants |
TRUE |
variant_call |
max_depth |
Int |
Maximum reads read at a position per input file for SAMtools mpileup before running iVar variants |
600000 |
variant_call |
min_bq |
Int |
Minimum mapping quality for an alignment to be used for SAMtools mpileup before running iVar variants |
0 |
variant_call |
min_depth |
Int |
Minimum read depth to call variants for iVar variants |
10 |
variant_call |
min_freq |
Float |
Minimum frequency threshold(0 - 1) to call variants for iVar variants |
0.6 |
variant_call |
min_qual |
Int |
Minimum quality threshold for sliding window to pass for iVar variants |
20 |
variant_call |
ref_gff |
String |
Path to the general feature format of the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/reference/GCF_009858895.2_ASM985889v3_genomic.gff |
variant_call |
ref_genome |
String |
Path to the reference genome within the staphb/ivar:1.2.2_artic20200528 Docker container |
/artic-ncov2019/primer_schemes/nCoV-2019/V3/nCoV-2019.reference.fasta |
version_capture |
timezone |
String |
User time zone in valid Unix TZ string (e.g. America/New_York) |
None |
Outputs¶
Download CSV: TheiaCoV_Illumina_SE_default_outputs.csv
Output Name |
Data Type |
Description |
|---|---|---|
aligned_bai |
File |
Index companion file to the bam file generated during the consensus assembly process |
aligned_bam |
File |
Primer-trimmed BAM file; generated during conensus assembly process |
assembly_fasta |
File |
Consensus genome assembly |
assembly_length_unambiguous |
Int |
Number of unambiguous basecalls within the SC2 consensus assembly |
assembly_mean_coverage |
Float |
Mean sequencing depth throughout the conesnsus assembly generated after performing primer trimming–calculated using the SAMtools coverage command |
assembly_method |
String |
Method employed to generate consensus assembly |
auspice_json |
File |
Auspice-compatable JSON output generated from NextClade analysis that includes the NextClade default samples for clade-typing and the single sample placed on this tree |
bbduk_docker |
String |
Docker image used to run BBDuk |
bwa_version |
String |
Version of BWA used to map read data to the reference genome |
consensus_flagstat |
File |
Output from the SAMtools flagstat command to assess quality of the alignment file (BAM) |
consensus_stats |
File |
Output from the SAMtools stats command to assess quality of the alignment file (BAM) |
fastqc_clean |
Int |
Number of reads after SeqyClean filtering as determined by FastQC |
fastqc_raw |
Int |
Number of reads after seqyclean filtering as determined by FastQC |
fastqc_version |
String |
Version of the FastQC software used for read QC analysis |
ivar_tsv |
File |
Variant descriptor file generated by iVar variants |
ivar_variant_version |
String |
Version of iVar for running the iVar variants command |
ivar_vcf |
File |
iVar tsv output converted to VCF format |
ivar_version_consensus |
String |
Version of iVar for running the iVar consensus command |
ivar_version_primtrim |
String |
Version of iVar for running the iVar trim command |
kraken_human |
Float |
Percent of human read data detected using the Kraken2 software |
kraken_report |
String |
Full Kraken report |
kraken_sc2 |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software |
kraken_version |
String |
Version of Kraken software used |
meanbaseq_trim |
Float |
Mean quality of the nucleotide basecalls aligned to the reference genome after primer trimming |
meanmapq_trim |
Float |
Mean quality of the mapped reads to the reference genome after primer trimming |
nextclade_aa_dels |
String |
Amino-acid deletions as detected by NextClade |
nextclade_aa_subs |
String |
Amino-acid substitutions as detected by NextClade |
nextclade_clade |
String |
NextClade clade designation |
nextclade_json |
File |
NexClade output in JSON file format |
nextclade_tsv |
File |
NextClade output in TSV file format |
nextclade_version |
String |
Version of NextClade software used |
number_Degenerate |
Int |
Number of degenerate basecalls within the consensus assembly |
number_N |
Int |
Number of fully ambiguous basecalls within the consensus assembly |
number_Total |
Int |
Total number of nucleotides within the consensus assembly |
pango_lineage |
String |
Pango lineage as detremined by Pangolin |
pango_lineage_report |
File |
Full Pango lineage report generated by Pangolin |
pangolin_assignment_version |
String |
Version of the pangolin software (e.g. PANGO or PUSHER) used for lineage asignment |
pangolin_conflicts |
String |
Number of lineage conflicts as deteremed by Pangolin |
pangolin_docker |
String |
Docker image used to run Pangolin |
pangolin_notes |
String |
Lineage notes as deteremined by Pangolin |
pangolin_versions |
String |
All Pangolin software and database version |
percent_reference_coverage |
Float |
Percent coverage of the reference genome after performing primer trimming; calculated as assembly_length_unambiguous / length of reference genome (SC2: 29,903) x 100 |
primer_bed_name |
String |
Name of the primer bed files used for primer trimming |
primer_trimmed_read_percent |
Float |
Percent of read data with primers trimmed as deteremined by iVar trim |
read1_clean |
File |
Forward read file after quality trimming and adapter removal |
samtools_version |
String |
Version of SAMtools used to sort and index the alignment file |
samtools_version_consensus |
String |
Version of SAMtools used to create the pileup before running iVar consensus |
samtools_version_primtrim |
String |
Version of SAMtools used to create the pileup before running iVar trim |
samtools_version_stats |
String |
Version of SAMtools used to assess quality of read mapping |
seq_platform |
String |
Description of the sequencing methodology used to generate the input read data |
theiacov_illumina_se_analysis_date |
String |
Date of analysis |
theiacov_illumina_se_version |
String |
Version of the Public Health Viral Genomics (PHVG) repository used |
trimmomatic_version |
String |
Version of Trimmomatic used |
vadr_alerts_list |
File |
File containing all of the fatal alerts as determined by VADR |
vadr_docker |
String |
Docker image used to run VADR |
vadr_num_alerts |
String |
Number of fatal alerts as determined by VADR |
TheiaCoV_ClearLabs¶
The TheiaCoV_ClearLabs workflow was written to process ClearLabs WGS read data for SARS-CoV-2 amplicon sequencing. Currently, Clear Labs sequencing is performed with the Artic V3 protocol. If alternative primer schemes such as the Qiaseq Primer Panel, the Swift Amplicon SARS-CoV-2 Panel and the Artic V4 Amplicon Sequencing Panel become avaialble on the platform, these data can can also be analysed with this workflow since the primer sequence coordinates of the PCR scheme utilized must be provided along with the raw Clear Labs read data must be provided in BED and FASTQ file formats, respectively.
Upon initiating a TheiaCoV_ClearLabs run, input ClearLabs read data provided for each sample will be processed to perform consensus genome assembly, infer the quality of both raw read data and the generated consensus genome, and assign SARS-CoV-2 lineage and clade types as outlined in the TheiaCoV_ClearLabs data workflow below.
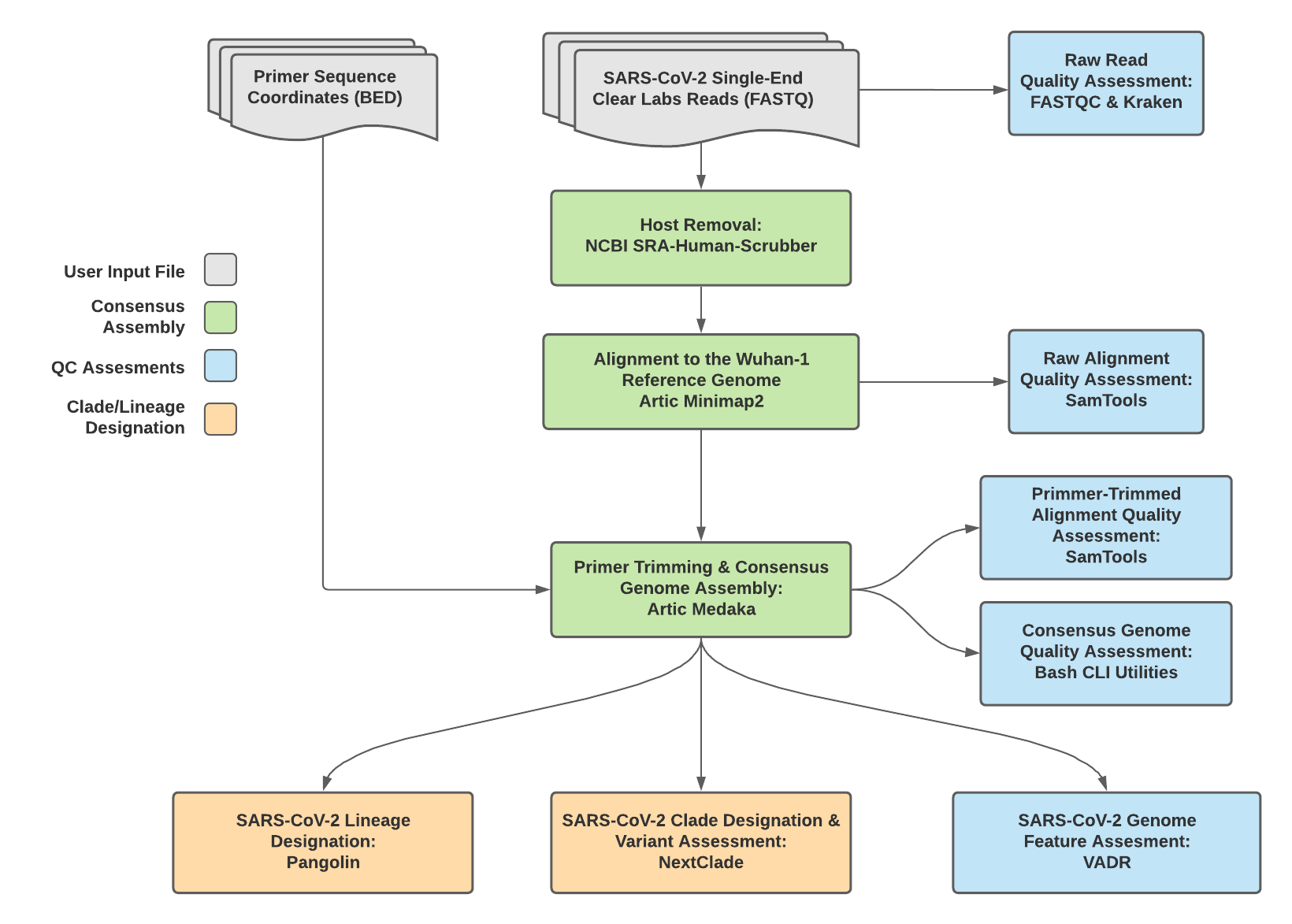
TheiaCoV_ClearLabs Data Workflow¶
Consensus genome assembly with the TheiaCoV_ClearLabs workflow is performed by first de-hosting read data with the NCBI SRA-Human-Scrubber tool then following the Artic nCoV-2019 novel coronavirs bioinformatics protocol <https://artic.network/ncov-2019/ncov2019-bioinformatics-sop.html>. Briefly, input reads are aligned to the Wuhan-1 reference genome with minimap2 to generate a Binary Alignment Mapping (BAM) file. Primer sequences are then removed from the BAM file and a consensus assembly file is generated using the Artic medaka command. This assembly is then used to assign lineage and clade designations with Pangolin and NextClade. NCBI’S VADR tool is also employed to screen for potentially errant features (e.g. erroneous frame-shift mutations) in the consensus assembly.
Note
Read-trimming is performed on raw read data generated on the ClearLabs instrument and thus not a required step in the TheiaCoV_ClearLabs workflow.
More information on required user inputs, optional user inputs, default tool parameters and the outputs generated by TheiaCoV_CLearLabs are outlined below.
Required User Inputs¶
Download CSV: TheiaCoV_ClearLabs_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
theiacov_clearlabs |
clear_lab_fastq |
File |
Clear Labs FASTQ read files |
theiacov_clearlabs |
primer_bed |
File |
Primer sequence coordinates of the PCR scheme utilized in BED file format |
theiacov_clearlabs |
samplename |
String |
Name of the sample being analyzed |
Optional User Inputs¶
Download CSV: TheiaCoV_ClearLabs_optional_inputs.csv
Task |
Variable Name |
Data Type |
Description |
Default |
|---|---|---|---|---|
consensus |
cpu |
Int |
CPU resources allocated to the Artric Medaka task runtime environment |
8 |
consensus |
docker |
String |
Docker tag used for running Medaka assemblyer |
staphb/artic-ncov2019:1.3.0 |
consensus |
medaka_model |
String |
Model for consensus genome assembly via Medaka |
r941_min_high_g360 |
fastqc_se_clean |
cpus |
Int |
CPU resources allocated to the FastQC task runtime environment for asessing clean read data |
|
fastqc_se_clean |
read1_name |
String |
Name of the sample being analyzed |
Inferred from the input read filefastqc_se_clean |
fastqc_se_raw |
cpus |
Int |
CPU resources allocated to the FastQC task runtime environment for asessing raw read data |
|
fastqc_se_raw |
read1_name |
String |
Name of the sample being analyzed |
Inferred from the input read file |
kraken2_dehosted |
cpus |
Int |
CPU resources allocated to the Kraken task runtime environment for asessing dehosted read data |
4 |
kraken2_dehosted |
kraken2_db |
String |
Path to the reference genome within the staphb/kraken2:2.0.8-beta_hv Docker container |
/kraken2-db |
kraken2_dehosted |
read2 |
File |
Optional input file for the Kraken task that is not applicable to this workflow |
None |
kraken2_raw |
cpus |
Int |
CPU resources allocated to the Kraken task runtime environment for asessing raw read data |
4 |
kraken2_raw |
kraken2_db |
String |
Path to the reference genome within the staphb/kraken2:2.0.8-beta_hv Docker container |
/kraken2-db |
kraken2_raw |
read2 |
File |
Optional input file for the Kraken task that is not applicable to this workflow |
None |
ncbi_scrub_se |
docker |
Docker tag used for running the NCBI SRA Human-Scruber tool |
gcr.io/ncbi-sys-gcr-public-research/sra-human-scrubber@sha256:b7dba71079344daea4ea3363e1a67fa54edb7ec65459d039669c68a66d38b140 |
|
nextclade_one_sample |
docker |
String |
Docker tag used for running NextClade |
neherlab/nextclade:0.14.2 |
nextclade_output_parser_one_sample |
docker |
String |
Docker tag used for parsing NextClade output |
python:slim |
pangolin3 |
docker |
String |
Docker tag used for running Pangolin |
staphb/pangolin:3.1.11-pangolearn-2021-08-24 |
pangolin3 |
inference_engine |
String |
pangolin inference engine for lineage designations (usher or pangolarn) |
usher |
pangolin3 |
min_length |
Int |
Minimum query length allowed for pangolin to attempt assignment |
10000 |
pangolin3 |
max_ambig |
Float |
Maximum proportion of Ns allowed for pangolin to attempt assignment |
0.5 |
theiacov_clearlabs |
nextclade_dataset_name |
String |
Nextclade organism dataset |
sars-cov-2 |
theiacov_clearlabs |
nextclade_dataset_reference |
String |
Nextclade reference genome |
MN908947 |
theiacov_clearlabs |
nextclade_dataset_tag |
Nextclade dataset tag |
2021-06-25T00:00:00Z |
|
theiacov_clearlabs |
normalise |
Int |
Value to normalize read counts |
200 |
theiacov_clearlabs |
seq_method |
String |
Description of the sequencing methodology used to generate the input read data |
ONT via Clear Labs WGS |
vadr |
docker |
String |
Docker tag used for running VADR |
staphb/vadr:1.2.1 |
vadr |
maxlen |
Int |
Maximum length for the fasta-trim-terminal-ambigs.pl VADR script |
30000 |
vadr |
minlen |
Int |
Minimum length subsequence to possibly replace Ns for the fasta-trim-terminal-ambigs.pl VADR script |
50 |
vadr |
skip_length |
Int |
Minimum assembly length (unambiguous) to run vadr |
10000 |
vadr |
vadr_opts |
String |
Options for the v-annotate.pl VADR script |
–glsearch -s -r –nomisc –mkey sarscov2 –alt_fail lowscore,fstukcnf,insertnn,deletinn –mdir /opt/vadr/vadr-models/ |
version_capture |
timezone |
String |
User time zone in valid Unix TZ string (e.g. America/New_York) |
None |
Outputs¶
Download CSV: TheiaCoV_ClearLabs_default_outputs.csv
Output Name |
Data Type |
Description |
|---|---|---|
aligned_bai |
File |
Index companion file to the bam file generated during the consensus assembly process |
aligned_bam |
File |
Primer-trimmed BAM file; generated during conensus assembly process |
artic_version |
String |
Version of the Artic software utilized for read trimming and conesnsus genome assembly |
assembly_fasta |
File |
Consensus genome assembly |
assembly_length_unambiguous |
Int |
Number of unambiguous basecalls within the SC2 consensus assembly |
assembly_mean_coverage |
Float |
Mean sequencing depth throughout the conesnsus assembly generated after performing primer trimming–calculated using the SAMtools coverage command |
assembly_method |
String |
Method employed to generate consensus assembly |
auspice_json |
File |
Auspice-compatable JSON output generated from NextClade analysis that includes the NextClade default samples for clade-typing and the single sample placed on this tree |
consensus_flagstat |
File |
Output from the SAMtools flagstat command to assess quality of the alignment file (BAM) |
consensus_stats |
File |
Output from the SAMtools stats command to assess quality of the alignment file (BAM) |
dehosted_reads |
File |
Dehosted reads; suggested read file for SRA submission |
fastqc_clean |
Int |
Number of reads after dehosting as determined by FastQC |
fastqc_raw |
Int |
Number of raw input reads as determined by FastQC |
fastqc_version |
String |
Version of the FastQC version used |
kraken_human |
Float |
Percent of human read data detected using the Kraken2 software |
kraken_human_dehosted |
Float |
Percent of human read data detected using the Kraken2 software after host removal |
kraken_report |
String |
Full Kraken report |
kraken_report_dehosted |
File |
Full Kraken report after host removal |
kraken_sc2 |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software |
kraken_sc2_dehosted |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software after host removal |
kraken_version |
String |
Version of Kraken software used |
meanbaseq_trim |
Float |
Mean quality of the nucleotide basecalls aligned to the reference genome after primer trimming |
meanmapq_trim |
Float |
Mean quality of the mapped reads to the reference genome after primer trimming |
nextclade_aa_dels |
String |
Amino-acid deletions as detected by NextClade |
nextclade_aa_subs |
String |
Amino-acid substitutions as detected by NextClade |
nextclade_clade |
String |
NextClade clade designation |
nextclade_json |
File |
NexClade output in JSON file format |
nextclade_tsv |
File |
NextClade output in TSV file format |
nextclade_version |
String |
Version of NextClade software used |
number_Degenerate |
Int |
Number of degenerate basecalls within the consensus assembly |
number_N |
Int |
Number of fully ambiguous basecalls within the consensus assembly |
number_Total |
Int |
Total number of nucleotides within the consensus assembly |
pango_lineage |
String |
Pango lineage as detremined by Pangolin |
pango_lineage_report |
File |
Full Pango lineage report generated by Pangolin |
pangolin_assignment_version |
String |
Version of the pangolin software (e.g. PANGO or PUSHER) used for lineage asignment |
pangolin_conflicts |
String |
Number of lineage conflicts as deteremed by Pangolin |
pangolin_docker |
String |
Docker image used to run Pangolin |
pangolin_notes |
String |
Lineage notes as deteremined by Pangolin |
pangolin_versions |
String |
All Pangolin software and database versions |
percent_reference_coverage |
Float |
Percent coverage of the reference genome after performing primer trimming; calculated as assembly_length_unambiguous / length of reference genome (SC2: 29,903) x 100 |
primer_bed_name |
String |
Name of the primer bed files used for primer trimming |
reads_dehosted |
File |
De-hosted read files |
samtools_version |
String |
Version of SAMtools used to sort and index the alignment file |
seq_platform |
String |
Description of the sequencing methodology used to generate the input read data |
theiacov_clearlabs_analysis_date |
String |
Date of analysis |
theiacov_clearlabs_version |
String |
Version of the Public Health Viral Genomics (PHVG) repository used |
vadr_alerts_list |
File |
File containing all of the fatal alerts as determined by VADR |
vadr_docker |
String |
Docker image used to run VADR |
vadr_num_alerts |
String |
Number of fatal alerts as determined by VADR |
variants_from_ref_vcf |
File |
Number of variants relative to the reference genome |
TheiaCoV_ONT¶
The TheiaCoV_ONT workflow was written to process basecalled and demultiplexed Oxford Nanopore Technology (ONT) read data. The most common read data analyzed by the TheiaCoV_ONT workflow are generated with the Artic V3 protocol. Alternative primer schemes such as the Qiaseq Primer Panel, the Swift Amplicon SARS-CoV-2 Panel and the Artic V4 Amplicon Sequencing Panel however, can also be analysed with this workflow since the primer sequence coordinates of the PCR scheme utilized must be provided along with the raw paired-end Illumina read data in BED and FASTQ file formats, respectively.
Upon initiating a TheiaCoV_ONT run, input ONT read data provided for each sample will be processed to perform consensus genome assembly, infer the quality of both raw read data and the generated consensus genome, and assign SARS-CoV-2 lineage and clade types as outlined in the TheiaCoV_ONT data workflow below.
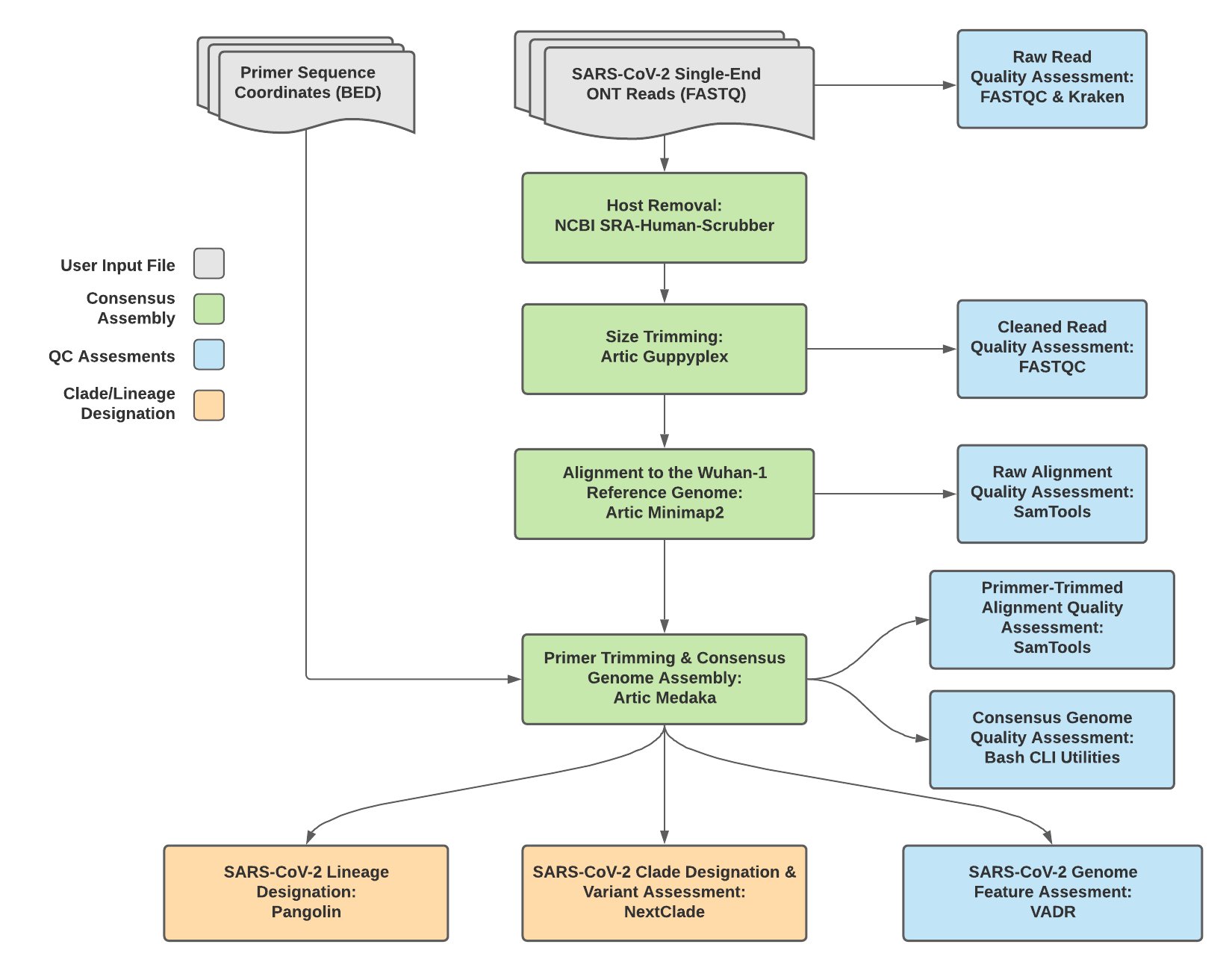
TheiaCoV_ONT Data Workflow¶
Consensus genome assembly with the TheiaCoV_ONT workflow is performed performed by first de-hosting read data with the NCBI SRA-Human-Scrubber tool then following then following Artic nCoV-2019 novel coronavirs bioinformatics protocol <https://artic.network/ncov-2019/ncov2019-bioinformatics-sop.html>. Briefly, input reads are filtered by size (min-length: 400bp; max-length: 700bp) with the Aritc guppyplex command. These size-selected read data are aligned to the Wuhan-1 reference genome with minimap2 to generate a Binary Alignment Mapping (BAM) file. Primer sequences are then removed from the BAM file and a consensus assembly file is generated using the Artic medaka command. This assembly is then used to assign lineage and clade designations with Pangolin and NextClade. NCBI’S VADR tool is also employed to screen for potentially errant features (e.g. erroneous frame-shift mutations) in the consensus assembly.
More information on required user inputs, optional user inputs, default tool parameters and the outputs generated by TheiaCoV_ONT are outlined below.
Required User Inputs¶
Download CSV: TheiaCoV_ONT_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
theiacov_ont |
demultiplexed_reads |
File |
Basecalled and demultiplexed ONT read data (single FASTQ file per sample) |
theiacov_ont |
primer_bed |
File |
Primer sequence coordinates of the PCR scheme utilized in BED file format |
theiacov_ont |
samplename |
String |
Name of the sample being analyzed |
Optional User Inputs¶
Download CSV: TheiaCoV_ONT_optional_inputs.csv
Task |
Variable Name |
Data Type |
Description |
Default |
|---|---|---|---|---|
consensus |
cpu |
Int |
CPU resources allocated to the Artric Medaka task runtime environment |
|
consensus |
docker |
String |
Docker tag used for running Medaka assemblyer |
staphb/artic-ncov2019:1.3.0 |
consensus |
medaka_model |
String |
Model for consensus genome assembly via Medaka |
r941_min_high_g360 |
fastqc_se_clean |
cpus |
Int |
CPU resources allocated to the FastQC task runtime environment for asessing size-selected read data |
2 |
fastqc_se_clean |
read1_name |
String |
Name of the sample being analyzed |
Inferred from the input read file |
fastqc_se_raw |
cpus |
Int |
CPU resources allocated to the FastQC task runtime environment for asessing raw read data |
|
fastqc_se_raw |
read1_name |
String |
Name of the sample being analyzed |
Inferred from the input read file |
kraken2_dehosted |
cpus |
Int |
CPU resources allocated to the Kraken task runtime environment for asessing dehosted read data |
4 |
kraken2_dehosted |
kraken2_db |
String |
Path to the reference genome within the staphb/kraken2:2.0.8-beta_hv Docker container |
/kraken2-db |
kraken2_dehosted |
read2 |
File |
Optional input file for the Kraken task that is not applicable to this workflow |
None |
kraken2_raw |
cpus |
Int |
CPU resources allocated to the Kraken task runtime environment for asessing raw read data |
4 |
kraken2_raw |
kraken2_db |
String |
Path to the reference genome within the staphb/kraken2:2.0.8-beta_hv Docker container |
/kraken2-db |
kraken2_raw |
read2 |
File |
Optional input file for the Kraken task that is not applicable to this workflow |
None |
ncbi_scrub_se |
docker |
Docker tag used for running the NCBI SRA Human-Scruber tool |
gcr.io/ncbi-sys-gcr-public-research/sra-human-scrubber@sha256:b7dba71079344daea4ea3363e1a67fa54edb7ec65459d039669c68a66d38b140 |
|
nextclade_one_sample |
docker |
String |
Docker tag used for running NextClade |
neherlab/nextclade:0.14.2 |
nextclade_output_parser_one_sample |
docker |
String |
Docker tag used for parsing NextClade output |
python:slim |
pangolin3 |
docker |
String |
Docker tag used for running Pangolin |
staphb/pangolin:3.1.11-pangolearn-2021-08-24 |
pangolin3 |
inference_engine |
String |
pangolin inference engine for lineage designations (usher or pangolarn) |
usher |
pangolin3 |
min_length |
Int |
Minimum query length allowed for pangolin to attempt assignment |
10000 |
pangolin3 |
max_ambig |
Float |
Maximum proportion of Ns allowed for pangolin to attempt assignment |
0.5 |
read_filtering |
cpu |
Int |
CPU resources allocated to the read filtering task (Artic guppypled) runtime environment |
8 |
read_filtering |
max_length |
Int |
Maximum sequence length |
700 |
read_filtering |
min_length |
Int |
Minimum sequence length |
400 |
read_filtering |
run_prefix |
String |
Run name |
artic_ncov2019 |
theiacov_ont |
nextclade_dataset_name |
String |
Nextclade organism dataset |
sars-cov-2 |
theiacov_ont |
nextclade_dataset_reference |
String |
Nextclade reference genome |
MN908947 |
theiacov_ont |
nextclade_dataset_tag |
Nextclade dataset tag |
2021-06-25T00:00:00Z |
|
theiacov_ont |
artic_primer_version |
String |
Version of the Artic PCR protocol used to generate input read data |
V3 |
theiacov_ont |
normalise |
Int |
Value to normalize read counts |
200 |
theiacov_ont |
seq_method |
String |
Description of the sequencing methodology used to generate the input read data |
ONT |
theiacov_ont |
pangolin_docker_image |
String |
Docker tag used for running Pangolin |
staphb/pangolin:2.4.2-pangolearn-2021-05-19 |
vadr |
docker |
String |
Docker tag used for running VADR |
staphb/vadr:1.2.1 |
vadr |
maxlen |
Int |
Maximum length for the fasta-trim-terminal-ambigs.pl VADR script |
30000 |
vadr |
minlen |
Int |
Minimum length subsequence to possibly replace Ns for the fasta-trim-terminal-ambigs.pl VADR script |
50 |
vadr |
vadr_opts |
String |
Options for the v-annotate.pl VADR script |
–glsearch -s -r –nomisc –mkey sarscov2 –alt_fail lowscore,fstukcnf,insertnn,deletinn –mdir /opt/vadr/vadr-models/ |
vadr |
skip_length |
Int |
Minimum assembly length (unambiguous) to run vadr |
10000 |
version_capture |
timezone |
String |
User time zone in valid Unix TZ string (e.g. America/New_York) |
None |
Outputs¶
Download CSV: TheiaCoV_ONT_default_outputs.csv
Output Name |
Data Type |
Description |
|---|---|---|
aligned_bai |
File |
Index companion file to the bam file generated during the consensus assembly process |
aligned_bam |
File |
Primer-trimmed BAM file; generated during conensus assembly process |
amp_coverage |
File |
Sequence coverage per amplicon |
artic_version |
String |
Version of the Artic software utilized for read trimming and conesnsus genome assembly |
assembly_fasta |
File |
Consensus genome assembly |
assembly_length_unambiguous |
Int |
Number of unambiguous basecalls within the SC2 consensus assembly |
assembly_mean_coverage |
Float |
Mean sequencing depth throughout the conesnsus assembly generated after performing primer trimming–calculated using the SAMtools coverage command |
assembly_method |
String |
Method employed to generate consensus assembly |
auspice_json |
File |
Auspice-compatable JSON output generated from NextClade analysis that includes the NextClade default samples for clade-typing and the single sample placed on this tree |
bedtools_version |
String |
bedtools version utilized when calculating amplicon read coverage |
consensus_flagstat |
File |
Output from the SAMtools flagstat command to assess quality of the alignment file (BAM) |
consensus_stats |
File |
Output from the SAMtools stats command to assess quality of the alignment file (BAM) |
dehosted_reads |
File |
Dehosted reads; suggested read file for SRA submission |
fastqc_clean |
Int |
Number of reads after size filttering and dehosting as determined by FastQC |
fastqc_raw |
Int |
Number of raw reads input reads as determined by FastQC |
fastqc_version |
String |
Version of the FastQC version used |
kraken_human |
Float |
Percent of human read data detected using the Kraken2 software |
kraken_human_dehosted |
Float |
Percent of human read data detected using the Kraken2 software after host removal |
kraken_report |
File |
Full Kraken report |
kraken_report_dehosted |
File |
Full Kraken report after host removal |
kraken_sc2 |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software |
kraken_sc2_dehosted |
Float |
Percent of SARS-CoV-2 read data detected using the Kraken2 software after host removal |
kraken_version |
String |
Version of Kraken software used |
meanbaseq_trim |
Float |
Mean quality of the nucleotide basecalls aligned to the reference genome after primer trimming |
meanmapq_trim |
Float |
Mean quality of the mapped reads to the reference genome after primer trimming |
nextclade_aa_dels |
String |
Amino-acid deletions as detected by NextClade |
nextclade_aa_subs |
String |
Amino-acid substitutions as detected by NextClade |
nextclade_clade |
String |
NextClade clade designation |
nextclade_json |
File |
NexClade output in JSON file format |
nextclade_tsv |
File |
NextClade output in TSV file format |
nextclade_version |
String |
Version of NextClade software used |
number_Degenerate |
Int |
Number of degenerate basecalls within the consensus assembly |
number_N |
Int |
Number of fully ambiguous basecalls within the consensus assembly |
number_Total |
Int |
Total number of nucleotides within the consensus assembly |
pango_lineage |
String |
Pango lineage as detremined by Pangolin |
pango_lineage_report |
File |
Full Pango lineage report generated by Pangolin |
pangolin_assignment_version |
String |
Version of the pangolin software (e.g. PANGO or PUSHER) used for lineage asignment |
pangolin_conflicts |
String |
Number of lineage conflicts as deteremed by Pangolin |
pangolin_docker |
String |
Docker image used to run Pangolin |
pangolin_notes |
String |
Lineage notes as deteremined by Pangolin |
pangolin_versions |
String |
All Pangolin software and database versions |
percent_reference_coverage |
Float |
Percent coverage of the reference genome after performing primer trimming; calculated as assembly_length_unambiguous / length of reference genome (SC2: 29,903) x 100 |
primer_bed_name |
String |
Name of the primer bed files used for primer trimming |
pangolin_versions |
String |
All Pangolin software and database versions |
reads_dehosted |
File |
De-hosted read files |
samtools_version |
String |
Version of SAMtools used to sort and index the alignment file |
seq_platform |
String |
Description of the sequencing methodology used to generate the input read data |
theiacov_ont_analysis_date |
String |
Date of analysis |
theiacov_ont_version |
String |
Version of the Public Health Viral Genomics (PHVG) repository used |
vadr_alerts_list |
File |
File containing all of the fatal alerts as determined by VADR |
vadr_docker |
String |
Docker image used to run VADR |
vadr_num_alerts |
String |
Number of fatal alerts as determined by VADR |
variants_from_ref_vcf |
File |
Number of variants relative to the reference genome |
TheiaCoV_FASTA¶
The TheiaCoV_FASTA workflow was written to process SARS-CoV-2 assembly files to infer the quality of the input assembly and assign SARS-CoV-2 lineage and clade types as outlined in the TheiaCoV_FASTA data workflow below.
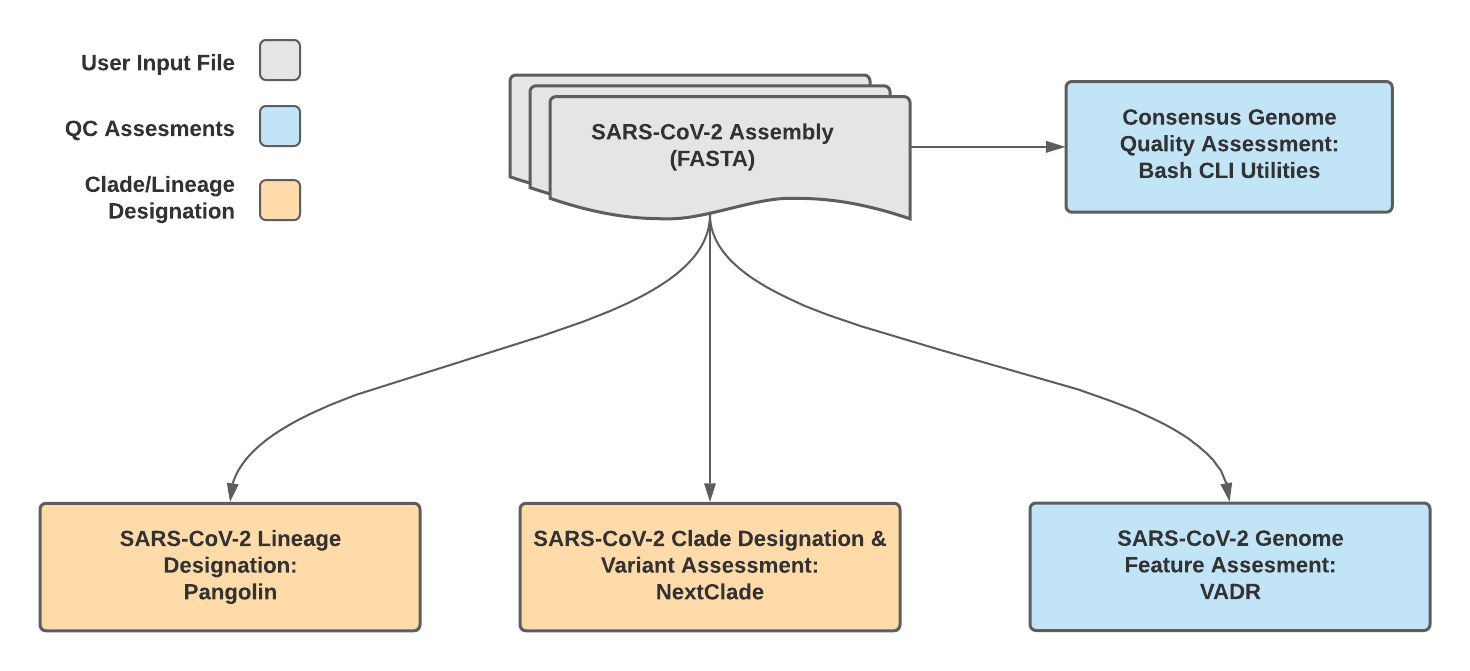
TheiaCoV_FASTA Data Workflow¶
The quality of input SARS-CoV-2 genome assemblies are assessed by the TheiaCoV_FASTA workflow using a series of bash shell scripts. Input assemblies are then used to assign lineage and clade designations with Pangolin and NextClade. NCBI’S VADR tool is also employed to screen for potentially errant features (e.g. erroneous frame-shift mutations) in the consensus assembly.
More information on required user inputs, optional user inputs, default tool parameters and the outputs generated by TheiaCoV_FASTA are outlined below.
Required User Inputs¶
Download CSV: TheiaCoV_FASTA_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
theiacov_fasta |
assembly_fasta |
File |
SARS-CoV-2 assemly file in fasta format |
theiacov_fasta |
input_assembly_method |
String |
Description of the method utilized to generate the input assembly fasta file; if unknown “NA” will be accepted |
theiacov_fasta |
samplename |
String |
Name of the sample being analyzed |
theiacov_fasta |
seq_method |
String |
Description of the sequencing method utilized to generate the raw sequencing data; if unknown “NA” will be accepted |
Optional User Inputs¶
Download CSV: TheiaCoV_FASTA_optional_inputs.csv
Task |
Variable Name |
Data Type |
Description |
Default |
|---|---|---|---|---|
nextclade_one_sample |
docker |
String |
Docker tag used for running NextClade |
neherlab/nextclade:0.14.2 |
nextclade_output_parser_one_sample |
docker |
String |
Docker tag used for parsing NextClade output |
python:slim |
pangolin3 |
docker |
String |
Docker tag used for running Pangolin |
staphb/pangolin:3.1.11-pangolearn-2021-08-24 |
pangolin3 |
inference_engine |
String |
pangolin inference engine for lineage designations (usher or pangolarn) |
usher |
pangolin3 |
max_ambig |
Float |
Maximum proportion of Ns allowed for pangolin to attempt assignment |
0.5 |
pangolin3 |
min_length |
Int |
Minimum query length allowed for pangolin to attempt assignment |
10000 |
titan_fasta |
nextclade_dataset_name |
String |
Nextclade organism dataset |
sars-cov-2 |
titan_fasta |
nextclade_dataset_reference |
String |
Nextclade reference genome |
MN908947 |
titan_fasta |
nextclade_dataset_tag |
Nextclade dataset tag |
2021-06-25T00:00:00Z |
|
vadr |
docker |
String |
Docker tag used for running VADR |
staphb/vadr:1.2.1 |
vadr |
maxlen |
Int |
Maximum length for the fasta-trim-terminal-ambigs.pl VADR script |
30000 |
vadr |
minlen |
Int |
Minimum length subsequence to possibly replace Ns for the fasta-trim-terminal-ambigs.pl VADR script |
50 |
vadr |
skip_length |
Int |
Minimum assembly length (unambiguous) to run vadr |
10000 |
vadr |
vadr_opts |
String |
Options for the v-annotate.pl VADR script |
–glsearch -s -r –nomisc –mkey sarscov2 –alt_fail lowscore,fstukcnf,insertnn,deletinn –mdir /opt/vadr/vadr-models/ |
version_capture |
timezone |
String |
User time zone in valid Unix TZ string (e.g. America/New_York) |
None |
Outputs¶
Download CSV: TheiaCoV_FASTA_default_outputs.csv
TheiaCoV Workflows for Genomic Epidemiology¶
Genomic Epidemiology, i.e. generating phylogenetic trees from a set of consensus assemblies (FASTA format) to track the spread and evolution of viruses on a local, national or global scale, has been an important methodological approach in the effort to mitigate disease transmission.
The TheiaCoV Genomic Epidemiology Series contains two seperate WDL workflows (TheiaCoV_Augur_Prep and TheiaCoV_Augur_Run) that process a set of viral genomic assemblies to generate phylogenetic trees (JSON format) and metadata files which can be used to assign epidemiological data to each assembly for subsequent analyses.
The two TheiaCoV workflows for genomic epidemiology must be run sequentially to first prepare the data for phylogenetic analysis and second to generate the phylogenetic trees. More information on the technical details of these processes and information on how to utilize and apply these workflows for public health investigations is available below.
Download CSV: TheiaCoV_Augur_Prep_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
prep_augur_metadata |
assembly |
File |
Assembly/consensus file (single FASTA file per sample) |
prep_augur_metadata |
collection_date |
String |
Collection date of the sample to be included in the analysis |
prep_augur_metadata |
iso_country |
String |
Country of the sample to be included in the analysis |
prep_augur_metadata |
iso_state |
String |
State of the sample to be included in the analysis |
prep_augur_metadata |
iso_continent |
String |
Continent of the sample to be included in the analysis |
prep_augur_metadata |
pango_lineage |
String |
Pango Lineage of the sample to be included in the analysis |
TheiaCoV_Augur_Prep¶
The TheiaCoV_Augur_Prep workflow was written to process consensus assemblies (FASTA format) and the associated metadata in preparation for running the TheiaCoV_Augur_Run. Input assemblies should be of similar quality (percent reference coverage, number of ambiguous bases, etc.). Inputs with highly discordant quality metrics may result in inaccurate inference of genetic relatedness.
Note
There must be some sequence diversity in the input set of assemblies to be analyzed. As a rule of thumb, the smaller the input set, the more sequence diversity will be required to make any sort of genomic inference. If a small (~10) set of viral genomic assemblies is used as the input then it may be necessary to add one significantly divergent assembly.
Upon initiating a TheiaCoV_Augur_Prep run, input assembly/consensus files and associated metadata will be used to produce the array of assembly/consensus files and the array of metadata files to be used as inputs for the TheiaCoV_Augur_Run workflow.
Metadata files are prepared with the Augur_Prep workflow by using BASH commands to first de-identify, and then to parse the headers of the input assembly files.
Required User Inputs¶
Download CSV: TheiaCoV_Augur_Prep_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
prep_augur_metadata |
assembly |
File |
Assembly/consensus file (single FASTA file per sample) |
prep_augur_metadata |
collection_date |
String |
Collection date of the sample to be included in the analysis |
prep_augur_metadata |
iso_country |
String |
Country of the sample to be included in the analysis |
prep_augur_metadata |
iso_state |
String |
State of the sample to be included in the analysis |
prep_augur_metadata |
iso_continent |
String |
Continent of the sample to be included in the analysis |
prep_augur_metadata |
pango_lineage |
String |
Pango Lineage of the sample to be included in the analysis |
TheiaCoV_Augur_Run¶
The TheiaCoV_Augur_Run workflow was written to process an array of assembly/consensus files (FASTA format) and and array of sample metadata files (TSV format) using a modified version of The Broad Institute’s sarscov2_nextstrain WDL workflow to create an Auspice JSON file; output from the modified sarscov2_nextstrain workflow will also be used to infer SNP distances and create a static PDF report.
Upon initiating a TheiaCoV_Augur_Run run, the input assembly/consensus file array and the associated metadata file array will be used to generate a JSON file that is compatible with phylogenetic tree building software. This JSON can then be used in Auspice or Nextstrain to view the phylogenetic tree. This phylogeneic tree can be used in genomic epidemiological analysis to visualize the genetic relatedness of a set of samples. The associated metadata can then be used to add context to the phylogenetic visualization.
Required User Inputs¶
Download CSV: TheiaCoV_Augur_Run_required_inputs.csv
Task |
Input Variable |
Data Type |
Description |
|---|---|---|---|
sarscov2_nextstrain |
assembly_fastas |
Array[File] |
An array of assembly/consensus files (FASTA) |
sarscov2_nextstrain |
sample_metadata_tsvs |
Array[File] |
An array of sample metadata files (TSV) |
sarscov2_nextstrain |
build_name |
String |
The name of the Augur build to be used in this analysis |